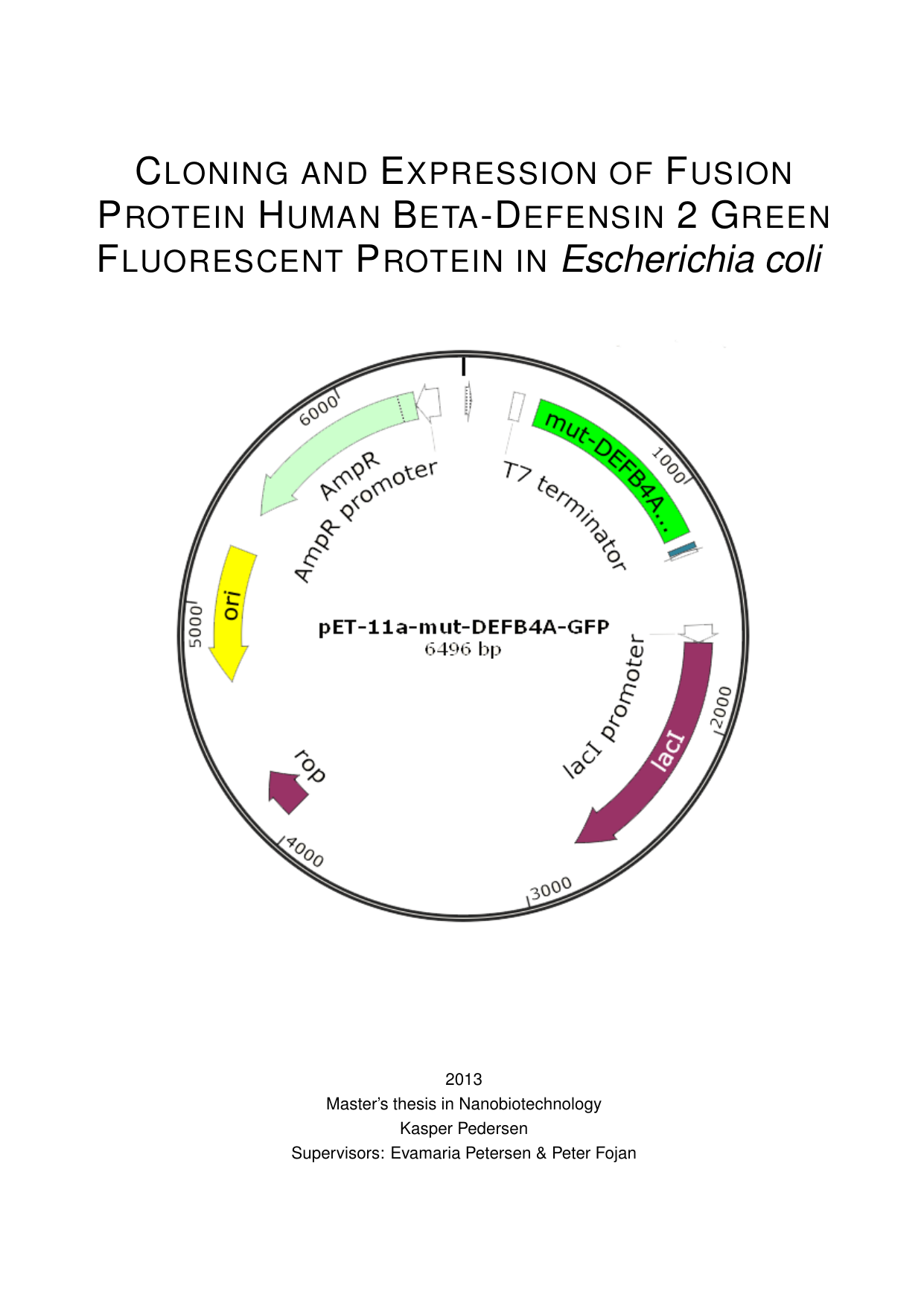
Cloning and Expression of Fusion Protein Human Beta-Defensin 2 Green Fluorescent Protein in Escherichia coli
Author
Pedersen, Kasper
Term
4. term
Education
Publication year
2013
Submitted on
2013-09-10
Pages
94
Abstract
In this master's thesis, we produced a soluble fusion protein linking HbD-2 to turboGFP in the bacterium E. coli (strain Origami 2(DE3)). A fusion protein joins two proteins so one can help track the other; here, turboGFP serves as a bright fluorescent tag. Fluorescence measurements showed that the turboGFP part was correctly folded and active. They also showed that cultures induced with IPTG (a chemical that switches on gene expression) contained much more fusion protein than uninduced controls. The expected size of the fusion protein was confirmed by SDS-PAGE, a gel method that separates proteins by size. To build the expression vector, we ligated DEFB4A-GFP into the pET-11a plasmid. Standard subcloning was complicated by an NdeI restriction site within the DEFB4A-GFP gene. A partial digestion with NdeI was attempted but was unsuccessful. We then introduced a targeted point mutation by PCR to change the NdeI recognition sequence from CATATG to CATTTG, thereby removing the internal NdeI site. The mutated gene was then digested with NdeI and BamHI and ligated into pET-11a to create pET-11a-mut-DEFB4A-GFP. Sequencing confirmed the construct and showed no mutations other than the intended point change.
I denne kandidatopgave fremstillede vi et opløseligt fusionsprotein, hvor HbD-2 er koblet til turboGFP, i bakterien E. coli (stammen Origami 2(DE3)). Et fusionsprotein er to proteiner, der er sat sammen, så det ene kan bruges til at følge det andet; her fungerer turboGFP som et stærkt fluorescerende mærke. Fluorescensmålinger viste, at turboGFP-delen var korrekt foldet og aktiv. De viste også, at kulturer induceret med IPTG (et kemikalie, der tænder for genekspression) producerede langt mere fusionsprotein end ikke-inducerede kontroller. Den forventede størrelse af fusionsproteinet blev bekræftet med SDS-PAGE, en gelmetode der adskiller proteiner efter størrelse. For at bygge expressionsvektoren ligerede vi DEFB4A-GFP ind i plasmidet pET-11a. Standard subkloning var vanskelig, fordi der fandtes et NdeI-restriktionssite inde i DEFB4A-GFP-genet. En delvis NdeI-fordøjelse blev forsøgt, men lykkedes ikke. Vi indførte derefter en målrettet punktmutation ved PCR, som ændrede NdeI-genkendelsessekvensen fra CATATG til CATTTG og dermed fjernede det interne NdeI-site. Det muterede gen kunne derefter fordøjes med NdeI og BamHI og ligeres ind i pET-11a for at danne pET-11a-mut-DEFB4A-GFP. Sekventering viste kun den tilsigtede punktmutation og ingen yderligere mutationer.
[This apstract has been rewritten with the help of AI based on the project's original abstract]
